The first report on the seahorse genome and the evolutionary status of its specialized morphology was recently published as a full and cover article in Nature on Dec 15th. The seahorse research project was directly led by Dr. Lin Qiang’ s group in the South China Sea Institute of Oceanology, CAS (SCSIO-CAS), with key joint contributions from Germany and Singapore research groups. This paper in Nature reveals the adaptive evolution of the seahorse to the sea shore and coral reef habitat (Nature Cover: evolution at a gallop). The Nature news titled “symbolic seahorse: the genome sequence of this unusual creature offers clues to its unique traits” highlights the evolution characteristics of the seahorse morphology and its reproductive mode that explained by the genes and the genome data.
The seahorses are widely distributed in the ocean throughout the world, and they are regarded as an important environmental indicator species in the marine ecosystem. Dr. Lin’ s group analyzed the whole genome data and found that the seahorse was the faster evolutionary species (Fig. 1). The seahorse has small numbers of olfactory receptor genes (OR) through comparing with other the ray-finned fishes. The seahorse also lacks enamel matrix protein-coding proline/glutamine- rich secretory calcium-binding phosphoprotein genes, which might lead to the loss of mineralized teeth. Comparative genomic analysis identified that the seahorse genome lost the great numbers of conserved noncoding elements (CNEs), which are involved in the development of the limbs and skeletal system. Male pregnancy is a unique developmental feature of seahorses and pipefishes. The astacin family of c6ast metalloprotease genes, which are responsible for hatching of embryos, have undergone tandem duplications given rise to six genes. Of the six duplicated genes in seahorse, five are highly expressed in the male brood pouch, suggesting that they may be involved in male pregnancy (Fig. 2).
The loss of pelvic fins in seahorse is associated with evolution of an armour-like covering of its body and gain of an elongated, flexible, substrate-gripping tail. A regulator gene, tbx4, is found lost in the seahorse genome. Knockout of tbx4 in zebrafish showed a ‘pelvic fin-loss’ phenotype similar to that of seahorses. The loss of the tbx4 gene may have a role in the loss of pelvic fins in seahorse (Fig. 3). These results provide important clues for elucidating the molecular mechanism of pelvic fin loss during fish evolution. It is of great significance to deepen the understanding of the biological characteristics of the seahorse and the evolution status of the teleost.
In conclusion, the research group led by Dr. Lin revealed the rapid evolution of seahorse genome and the mechanism of male pregnancy. These results will provide new insights into the relationships between the genes and adaptive evolution to the environment for some marine fishes. The research work was funded by the National Science Fund for Excellent Young Scholars (41322038), the Strategic Priority Research Program of the Chinese Academy of Sciences (XDA13020103), Youth Foundation of National High Technology Research and Development Program (2015AA020909) and other funding.
Article: http://www.nature.com/nature/journal/v540/n7633/full/nature20595.html;
Nature news: http://www.nature.com/news/sequence-reveals-genes-behind-bizarre-sea-horse-traits-1.21149

Nature Cover (Seahorse: evolution at a gallop)
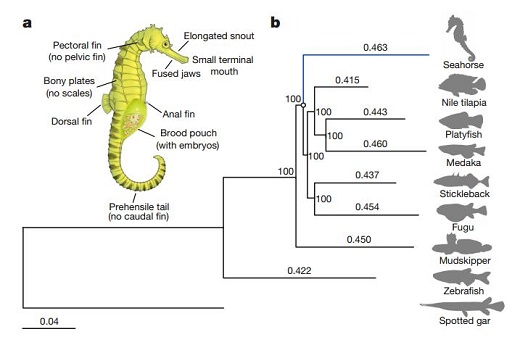
Figure 1. The morphology and the evolution rate of seahorse
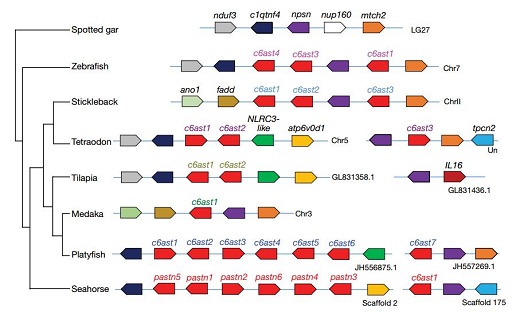
Figure 2. The duplication of pastn genes in seahorse
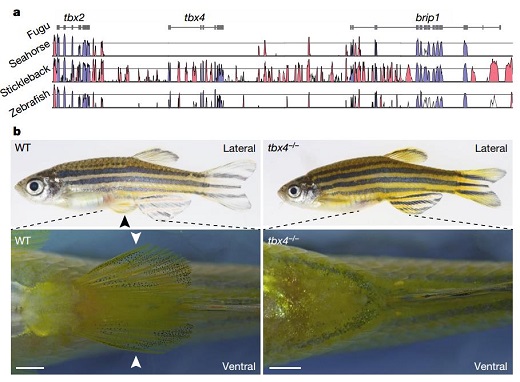
Figure 3. The gene knockout of tbx4 in zebrafish and the phenotype of the pelvic fins loss
